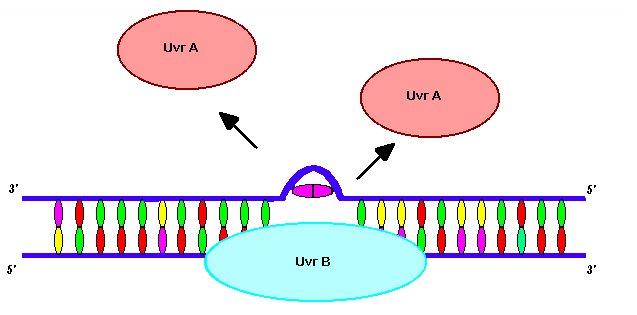
After TRCF has removed the RNA polymerase complex the NER cascade is free to begin. The UvrA subunits first dissociate from the UvrA 2 B complex leaving only UvrB. UvrB produces a severe bend (not shown) in the DNA at the damaged site. This bend helps with the dissociation of UvrA and site recognition by UvrC (Snustad and Simmons, 2000).

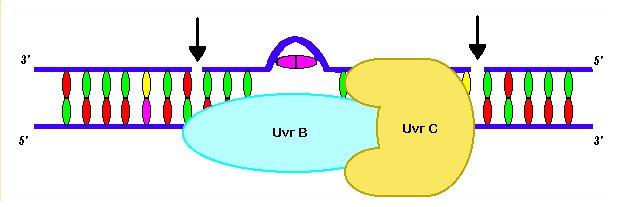
The next enzyme in the cascade is UvrC, which recognizes the bent UvrB-DNA complex. When bound together UvrB and UvrC for the UvrBC complex, which has endonulcease activity. The complex nicks the DNA at the 4th phosophodiester bond on the 3’ side and the 8th phosphodiester bond on the 5’ side of the lesion (Selby and Sancar, 1993). The cut creates a DNA oligomer that is approximately 12 nucleotides long (Snustad and Simmons, 2000).
After the UvrBC complex has cleaved the damaged strand, UvrD, also known as DNA helicase II, unpairs the DNA oligomer from the non-damaged strand. UvrC also departs at this stage (Snustad and Simmons, 2000).
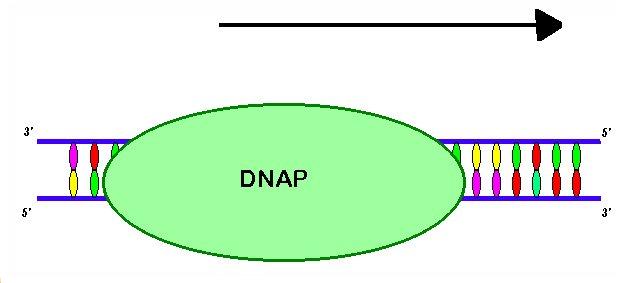
Finally, the gap left by the excised oligomer is filled in with new base pairs by DNA polymerase I and the nicks left in the repaired strand are sealed by DNA ligase. DNA polymerase I removes UvrB from the undamaged strand allowing DNA to resume its normal confirmation. After repair is complete, transcription will be initiated at the promoter region for the gene (Sunstad and Simmons, 2000). Remember that prokaryotic transcription-coupled repair that the RNA complex was removed and its transcript degraded. In eukaryotes, the repair of the DNA simply allows the stalled RNA complex to continue through instead of starting from scratch (Ahn and Grossman, 1996).